
YT-SOFTWARE
YT-SOFTWARE® offers a complete imaging toolbox
Image acquisition, device controlling as well as image and data analysis software for SYNENTEC's portfolio of automated imaging systems. YT-SOFTWARE® supports over 64 pre-configured imaging apps - ready to use - that are continuously extended and provide all applications for your high-throughput R&D processes in CLD and science.
Discover all the advantages
Image Cytometry – From Cells to Numbers
Turn your images into meaningful numbers
Export capabilities
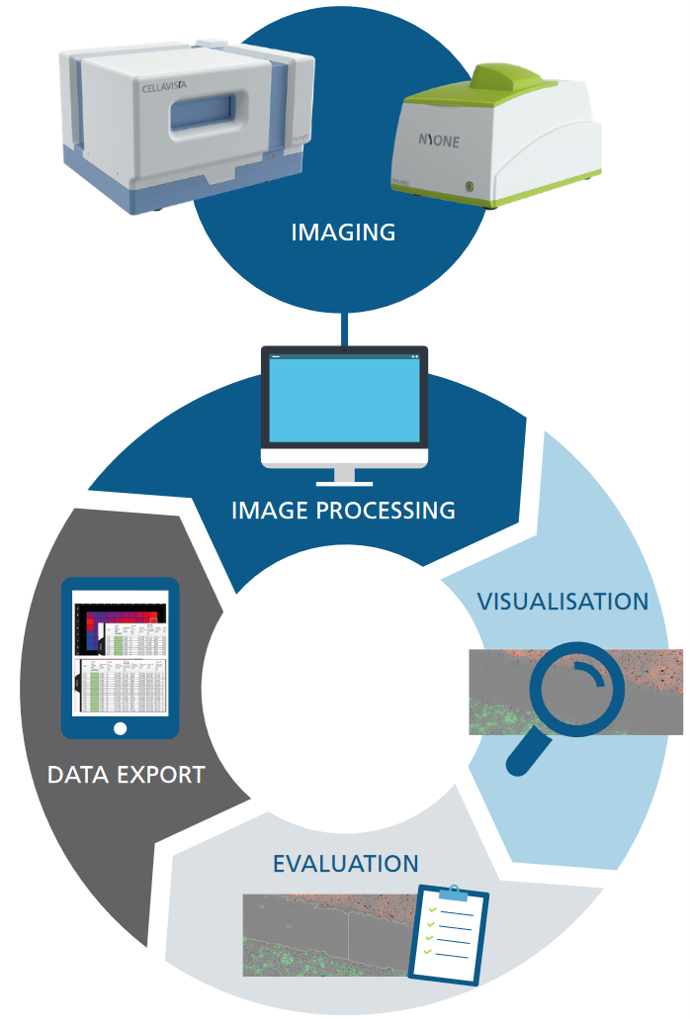
The YT-SOFTWARE® offers a complete toolbox: starting from experimental set-up and automatic high throughput image acquisition over precise image analysis up towards the storage, handling and processing of data for documentation and presentation purposes. And the high-resolution, stored images mean your results can be tracked over years to come.
Exportable Settings
- Experimental settings
- Image processing settings
Exportable Data
- Result table
- High content object data
- Plate heat map
- Growth curves / Time charts
- Histogram
- Scatter plot
Exportable Images
- Clone galery for SCC-report
- Plate overview
- Well channels & well overlay
- All taken Raw-images as ZIP-container
- Add brightness, contrast and normalization settings to image
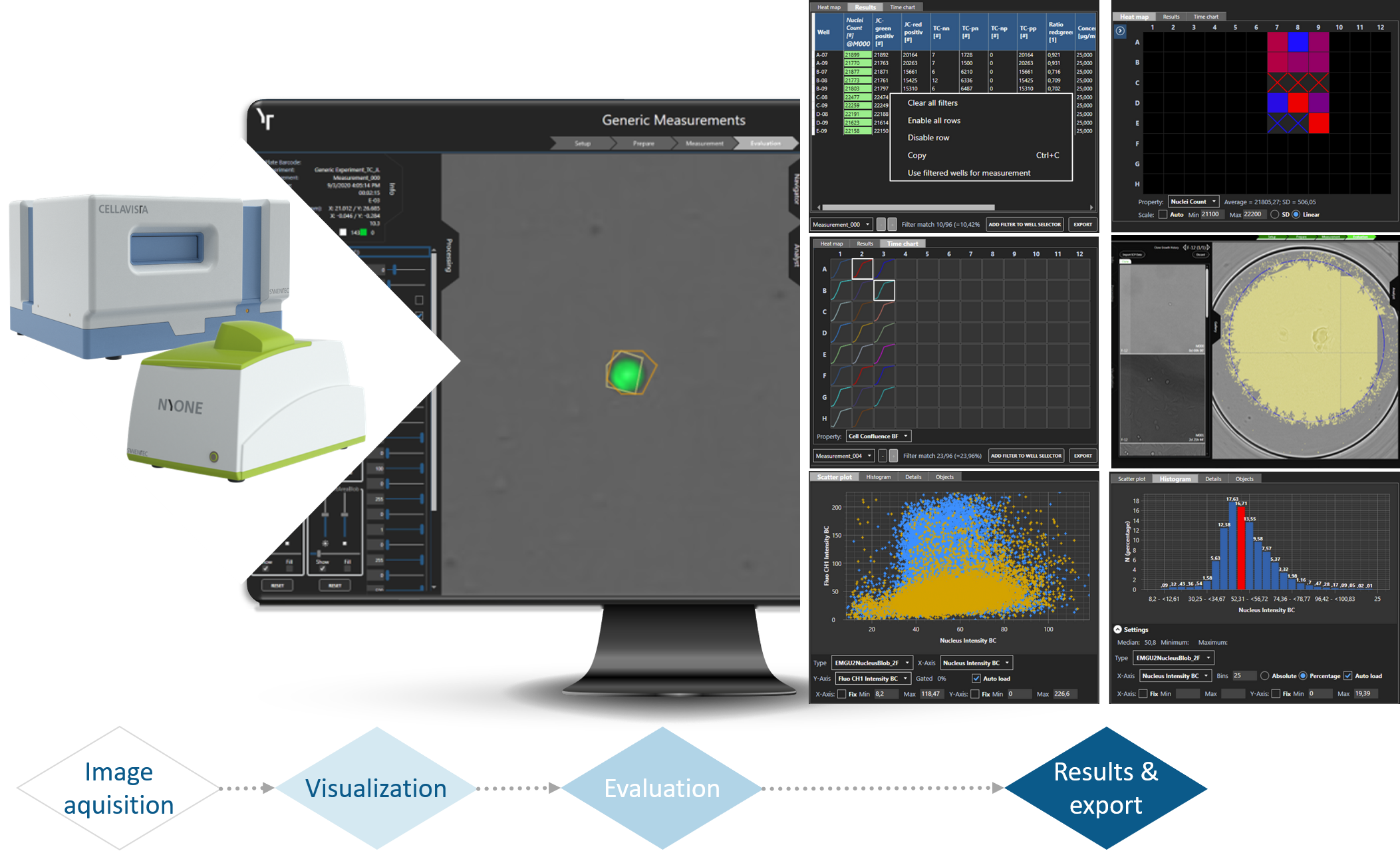
Data presentation in YT-SOFTWARE
DO YOU WANT TO KNOW MORE?
We know that time is an increasingly scarce resource, even in the lab. That's why we've thought your problem through and have everything ready for a complete one-handed solution.



